Explanation and analysis¶
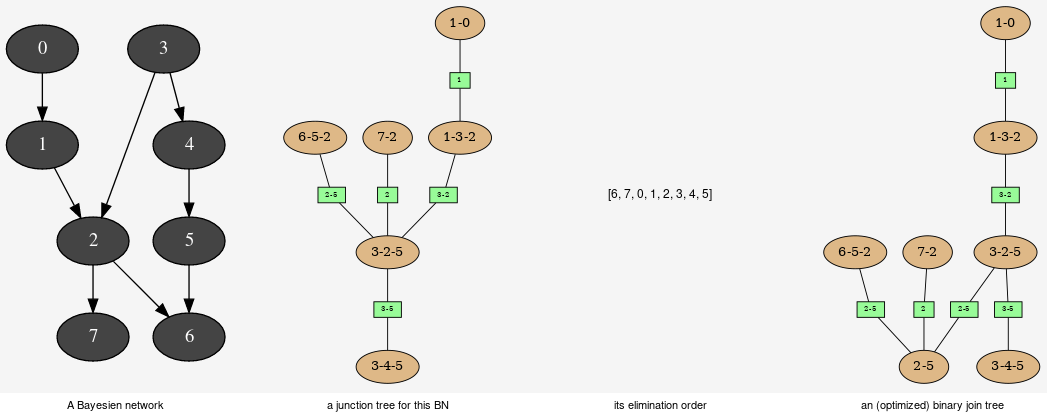
This tools aimed to provide some different views on the Bayesian network in order to explore its qualitative and/or quantitave behaviours.
-
class
pyAgrum.JunctionTreeGenerator¶ JunctionTreeGenerator is use to generate junction tree or binary junction tree from bayesian networks.
- JunctionTreeGenerator() -> JunctionTreeGenerator
- default constructor
-
binaryJoinTree(JunctionTreeGenerator self, UndiGraph g, PyObject * partial_order=None)¶ binaryJoinTree(JunctionTreeGenerator self, DAG dag, PyObject * partial_order=None) -> CliqueGraph binaryJoinTree(JunctionTreeGenerator self, BayesNet bn, PyObject * partial_order=None) -> CliqueGraph
Computes the binary joint tree for its parameters. If the first parameter is a graph, the heurisitcs assume that all the node have the same domain size (2). If given, the heuristic takes into account the partial order for its elimination order.
Parameters: - g (pyAgrum.UndiGraph) – a undirected graph
- dag (pyAgrum.DAG) – a dag
- bn (pyAgrum.BayesNet) – a BayesianNetwork
- partial_order (List[List[int]]) – a partial order among the nodeIDs
Returns: the current binary joint tree
Return type:
-
eliminationOrder(JunctionTreeGenerator self, UndiGraph g, PyObject * partial_order=None)¶ eliminationOrder(JunctionTreeGenerator self, DAG dag, PyObject * partial_order=None) -> PyObject eliminationOrder(JunctionTreeGenerator self, BayesNet bn, PyObject * partial_order=None) -> PyObject
Computes the elimination for its parameters. If the first parameter is a graph, the heurisitcs assume that all the node have the same domain size (2). If given, the heuristic takes into account the partial order for its elimination order.
Parameters: - g (pyAgrum.UndiGraph) – a undirected graph
- dag (pyAgrum.DAG) – a dag
- bn (pyAgrum.BayesNet) – a BayesianNetwork
- partial_order (List[List[int]]) – a partial order among the nodeIDs
Returns: the current elimination order.
Return type:
-
junctionTree(JunctionTreeGenerator self, UndiGraph g, PyObject * partial_order=None)¶ junctionTree(JunctionTreeGenerator self, DAG dag, PyObject * partial_order=None) -> CliqueGraph junctionTree(JunctionTreeGenerator self, BayesNet bn, PyObject * partial_order=None) -> CliqueGraph junctionTree(JunctionTreeGenerator self, MarkovNet mn, PyObject * partial_order=None) -> CliqueGraph
Computes the junction tree for its parameters. If the first parameter is a graph, the heurisitcs assume that all the node have the same domain size (2). If given, the heuristic takes into account the partial order for its elimination order.
Parameters: - g (pyAgrum.UndiGraph) – a undirected graph
- dag (pyAgrum.DAG) – a dag
- bn (pyAgrum.BayesNet) – a BayesianNetwork
- partial_order (List[List[int]]) – a partial order among the nodeIDs
Returns: the current junction tree.
Return type:
-
class
pyAgrum.EssentialGraph(*args)¶ Proxy of C++ pyAgrum.EssentialGraph class.
-
arcs(EssentialGraph self)¶ Returns: The lisf of arcs in the EssentialGraph Return type: list
-
children(EssentialGraph self, int id)¶ Parameters: id (int) – the id of the parent Returns: the set of all the children Return type: Set
-
connectedComponents()¶ connected components from a graph/BN
Compute the connected components of a pyAgrum’s graph or Bayesian Network (more generally an object that has nodes, children/parents or neighbours methods)
The firstly visited node for each component is called a ‘root’ and is used as a key for the component. This root has been arbitrarily chosen during the algorithm.
Returns: dict of connected components (as set of nodeIds (int)) with a nodeId (root) of each component as key. Return type: dict(int,Set[int])
-
edges(EssentialGraph self)¶ Returns: the list of the edges Return type: List
-
ids()¶ Deprecated method in pyAgrum>0.12.0. See nodes instead.
-
mixedGraph(EssentialGraph self)¶ Returns: the mixed graph Return type: pyAgrum.MixedGraph
-
neighbours(EssentialGraph self, int id)¶ Parameters: id (int) – the id of the checked node Returns: The set of edges adjacent to the given node Return type: Set
-
nodes(EssentialGraph self)¶
-
parents(EssentialGraph self, int id)¶ Parameters: id – The id of the child node Returns: the set of the parents ids. Return type: Set
-
size(EssentialGraph self)¶ Returns: the number of nodes in the graph Return type: int
-
sizeArcs(EssentialGraph self)¶ Returns: the number of arcs in the graph Return type: int
-
sizeEdges(EssentialGraph self)¶ Returns: the number of edges in the graph Return type: int
-
sizeNodes(EssentialGraph self)¶ Returns: the number of nodes in the graph Return type: int
-
skeleton(EssentialGraph self)¶
-
toDot(EssentialGraph self)¶ Returns: a friendly display of the graph in DOT format Return type: str
-
-
class
pyAgrum.MarkovBlanket(*args)¶ Proxy of C++ pyAgrum.MarkovBlanket class.
-
arcs(MarkovBlanket self)¶ Returns: the list of the arcs Return type: List
-
children(MarkovBlanket self, int id)¶ Parameters: id (int) – the id of the parent Returns: the set of all the children Return type: Set
-
connectedComponents()¶ connected components from a graph/BN
Compute the connected components of a pyAgrum’s graph or Bayesian Network (more generally an object that has nodes, children/parents or neighbours methods)
The firstly visited node for each component is called a ‘root’ and is used as a key for the component. This root has been arbitrarily chosen during the algorithm.
Returns: dict of connected components (as set of nodeIds (int)) with a nodeId (root) of each component as key. Return type: dict(int,Set[int])
-
dag(MarkovBlanket self)¶ Returns: a copy of the DAG Return type: pyAgrum.DAG
-
hasSameStructure(MarkovBlanket self, DAGmodel other)¶ Parameters: pyAgrum.DAGmodel – a direct acyclic model Returns: True if all the named node are the same and all the named arcs are the same Return type: bool
-
nodes(MarkovBlanket self)¶ Returns: the set of ids Return type: set
-
parents(MarkovBlanket self, int id)¶ Parameters: id – The id of the child node Returns: the set of the parents ids. Return type: Set
-
size(MarkovBlanket self)¶ Returns: the number of nodes in the graph Return type: int
-
sizeArcs(MarkovBlanket self)¶ Returns: the number of arcs in the graph Return type: int
-
sizeNodes(MarkovBlanket self)¶ Returns: the number of nodes in the graph Return type: int
-
toDot(MarkovBlanket self)¶ Returns: a friendly display of the graph in DOT format Return type: str
-